The Superorganism Revolution
By Robert Dorit
The bacteria living on and in us are challenging paradigms in community ecology.
The bacteria living on and in us are challenging paradigms in community ecology.

DOI: 10.1511/2014.110.330
In 1676, Antoni van Leeuwenhoek—a Dutch draper and amateur naturalist—peered through a microscope of his own design and described a world that would be misunderstood for the next 300 years. What he saw resulted in the first known description of bacteria, living beings which “were … so small in my eye…that if 100 of them lay one by another, they would not equal the length of a grain of course [sic] Sand....” No one had seen a living thing this small before.
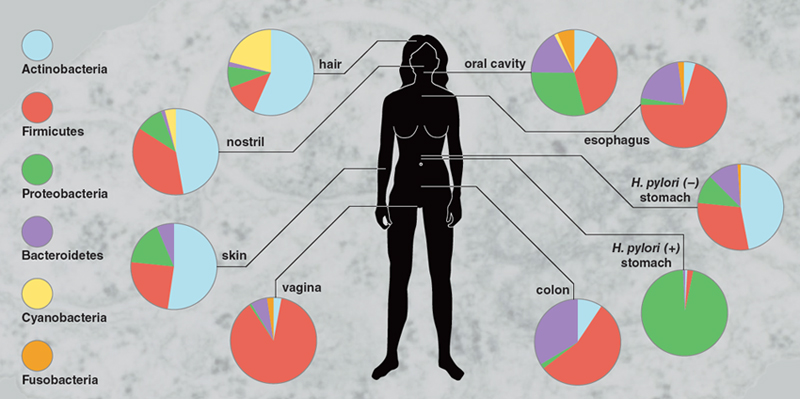
Illustration by Tom Dunne, adapted from I. Cho and M. Blaser, Nature Reviews Genetics 13:260.
But thanks to Robert Hooke, then Curator of Experiments of the Royal Society, incredulity gave way to acceptance. Hooke was the author in 1665 of Micrographia, the first illustrated account of microscopic observations and a likely inspiration for van Leeuwenhoek’s undertakings. In 1667, Hooke would confirm the observations of his Dutch colleague. The draper whom he called “Ingenious and Inquisitive” had seen and provided the first description of bacteria.
The existence of a living world beyond the reach of our senses is less mysterious today, but the medical profession’s attitude towards our bacterial associates has, until recently, oscillated between benign neglect and suspicious distrust.
Science is just starting to grasp the sheer abundance and diversity of bacterial life present on and in our bodies. More important, we realize that these bacteria are not simply squatters or unavoidable hitchhikers picked up as we move through a world crowded with microbes. Rather, they influence our health, digestion, metabolism, and response to medicines, not to mention our survival and evolution. The discovery of the human microbiome, the collection of microbial ecosystems that colonize virtually every external and internal body surface, has forever changed how we see ourselves. These bacteria shape our biology from birth to the grave. They are part of us.
A new subfield in the life sciences, somewhat clumsily tagged as microbiomics, has emerged in the 21st century to study the incredible 100 trillion bacteria that make us humans what we are. At the same time, the discovery that we host multiple microbial ecosystems has led to the resurgence of principles of ecology set down over the past century. Much as molecular biologists, facing unintelligible sequence data, rediscovered the importance of evolution in the 1990s, microbiologists are suddenly attuned to ecological methods and principles—developed in a variety of ecosystems ranging from tropical rainforests to the Pacific intertidal.
In the last half of the 20th century, mathematical theory, modeling, observation, and experiments gave rise to a mature and relatively unified theory that accounted for the distribution, abundance, and stability of terrestrial and aquatic ecosystems—composed primarily of readily visible organisms. In the new microbiome paradigm, the landscape inside the human body supports an ecosystem as complex as a rainforest or coral reef, and researchers study it in much the same way. Freed, thanks to sequencing, from the constraint of culturing bacteria to identify them, initial forays into the ecology of the microbiome have been primarily descriptive.
Scientists also need to probe for the mechanisms behind these descriptions—including the extent to which existing ecological “rules” apply at the microbiome scale. The discovery of the human microbiome, an interlocking set of functional microbial communities, provides the opportunity to test the universality of ecological models. Much as the discovery of bacteria forced a reevaluation of the scale at which life could be found, the sizes and time scales relevant to the microbiome are fundamentally different from anything studied before, raising the possibility that ecological principles will also need radical revision.
The human microbiome is not a single continuous ecosystem. Instead, it has evolved and differentiated to occupy five reasonably distinct body habitats: the skin, the nose, the mouth, the lower gastrointestinal tract, and the vagina. Each of these major habitats is, in turn, further subdivided. Thus, the microbial community inhabiting the lining of the cheeks differs significantly from the one that inhabits dental plaque.
These are early days in the study of the microbiome, and biologists are still not sure how many different bacteria make up each of these habitats. Nonetheless, early returns are quite staggering: The lower gut, the most diverse of our bacterial ecosystems, may house more than 30,000 different strains. The oral cavity ranks a close second, and—as your mother warned you—the area behind your ear is not far behind. These levels of diversity amaze, and they exceed even the most generous estimates of the diversity in tropical rainforests, where perhaps 15,000 different species might be found in an acre of undisturbed habitat.
These initial estimates are coarse and only begin to reflect the complexities of the microbiome. A more subtle analysis might focus on the number of bacterial cells of each species making up these microbial communities. Here too, a general pattern emerges. A small handful of species are enormously abundant, whereas the rest of the citizens occupy the surprisingly long tail of the distribution. This pattern has both methodological and ecological implications. Methodologically, microbiome studies require deep and concerted sampling to construct a true picture of diversity. Rare species are, by definition, easier to miss, but sampling only what is common can lead to misleading conclusions. The ecological implications of this long tail are no less profound: “Rare” does not mean “unimportant.” Which species are most abundant differs within the same individual from one environment to the next, changes over an individual’s lifetime, and varies within a particular body part from one individual to the next.
The notion that greater ecosystem diversity results in more stable ecosystems is certainly beguiling. More species may imply greater redundancy, connectivity, and capacity to absorb perturbations. But some ecologists have argued that more diverse ecosystems could also prove more fragile, their very complexity acting to magnify small disturbances into large effects.
The stability and resilience of the microbiome are now being actively studied, and the picture remains murky. On the one hand, the enormous diversity and apparent functional equivalence of a number of species mean that the loss of any given member of the community can be quickly compensated for by survivors. On the other hand, certain perturbations are dramatic and long-lasting in their consequences.
Over the past five years, the Human Microbiome Project—a consortium of laboratories—has been laying the groundwork for understanding these dynamics. They demonstrated that antibiotic use severely disrupts the microbiome, causing extensive collateral damage. The indiscriminate killing of nonpathogenic members of the microbiome makes it easier for pathogens to invade otherwise stable, occupied environments. As a result, pathogens that would not have a real chance of establishing themselves, most notably the aptly named Clostridium difficile, can run the table. Over the longer term, repeated antibiotic use may prevent the microbiome from ever recovering its original composition. Instead, such perturbed ecosystems may settle on a new composition that includes different species, many of them resistant to antibiotic treatment.
Evidence for succession—the replacement of early colonists by later arrivals, leading to a predictable pattern for each ecosystem—is abundant in both terrestrial and aquatic communities. For example, after the eruption of Mount Saint Helens in May of 1980, early colonists, dispersed by wind and able to exploit the barren volcanic soils, prepared the terrain for later species to come, ensuring their own disappearance in the process.
The human microbiome, too, undergoes succession, particularly in the first 24 to 36 months of life. We now understand, for example, that the passage through the birth canal seeds the newborn’s microbiome. Infants delivered by cesarean section, in contrast, exhibit a distinct microbiome that more closely resembles the composition of the mother’s skin. These initial colonization events leave a clear trace in the infant’s gut microbiome, a trace that can last months, years, and in some cases a lifetime.
The infant microbiome transitions in a somewhat predictable way to one of many possible stable configurations, but as elsewhere in biology, history matters. A number of factors, including maternal health prior to delivery, diet (breastfeeding versus formula feeding), and subsequent exposure to other bacteria, shape this progression of the microbiome to its adult form. The minuet between the host and the microbiome is ongoing: The evenness and composition of the gut microbiome, for instance, changes on a daily basis. Because the microbiome is clearly linked to the maturation and regulation of the human immune system, the composition of the microbiome is simultaneously shaping and being shaped by the biology of the host.
As a general rule in ecology, a few species are very abundant, but most species in a community are relatively or extremely sparse. The shape of the curve ranking species by their abundance tends to be mathematically well behaved, approximating lognormal or geometric distributions. It may be too soon to tell if the distribution of species’ abundances in the microbiome will adhere to this established ecological rule.
As I suggested earlier, the tail of the microbiome’s rank–abundance distribution is unexpectedly long. Just a few cells, sometimes fewer than 100 per species, represent hundreds of bacterial species. Given the dynamics of the microbiome, of course, membership in this tail may be transient. Under the right conditions, bacteria are capable of rapid population growth, and a species that appears rare in one snapshot of the microbiome could be present in high numbers a few hours later.
That caveat notwithstanding, the long tail is now a recurrent observation in the majority of well-studied microbiomes. Ecologists have long understood that the relevance of a species to the overall functioning of an ecosystem is not always reflected in its abundance. Indeed, certain species can be rare but are still critical to the stability and diversity of a community. Such keystone species, as their name suggests, engender the collapse of the entire ecosystem when they are removed. Although certain preliminary studies have sought to identify keystone microbiome species, a different and provocative alternative emerging from the work of the Human Microbiome Project is that these ecosystems may instead harbor keystone roles.
In this formulation, certain metabolic tasks must be performed in a functioning microbiome. Carbon and energy must be obtained, electrons properly handled, and waste products eliminated. But the specific identity of the species performing these requisite tasks may matter little. This interchangeability of species might explain how every human being can host a different, customized microbiome. Marked differences between one human microbiome and the next suggest that no single bacterial species must always be present in the gut—or in any other body environment—to ensure a working microbiome.
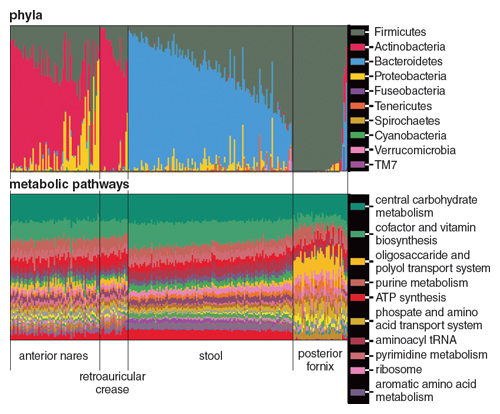
Illustration adapted by Tom Dunne, from The Human Microbiome Project, Nature 486:207.
If we focus not on species identity but on functional roles, the differences between environments within our body, and the differences from one microbiome to the next, begin to disappear (see graph above). Individual bacterial species matter only insofar as they can provide a specific ecological service to the microbiome, which, in turn, supplies every one of its component species with metabolites, nutrients, and raw materials. This organization ensures, among other things, efficient removal of potentially toxic by-products of metabolism, and provides a homeostatic environment for bacterial growth.
This unusual relationship between individual microbial species and the microbiome community to which they belong has profound implications. In the bacterial world, this relationship is stranger still. As we have learned over the past two decades, bacteria engage in extensive horizontal gene transfer, where functioning modules of genetic information are actively or passively exchanged across species boundaries. This trading of genetic information obeys neither lines of descent nor rules of shared ancestry. To a surprising extent, it enables bacteria to acquire functional genetic information in response to environmental change without having to evolve it de novo.
Horizontal gene transfer has attracted our attention because it underlies the rapid spread of antibiotic resistance and accounts for the acquisition of a pathogenic lifestyle in a number of species. But it may also cement the ability of the microbiome to respond to various perturbations, including changes in host health. Effectively, the microbiome is the creator and custodian of a repository of evolved information, potentially shared by every member of the microbiome.
Most of the genetic information inside you is not really “you”: The collective microbial genome in our gut may include 100-fold more genetic information than what can be found in our own eukaryotic cells. The information contained in this community repository is in constant motion, even across vast phylogenetic gulfs. This library—the result of billions of natural experiments that have been unfolding over the past 3 billion years—is a real and coherent evolving entity and may be the key to microbiome persistence. In contrast to the traditional focus on the individual organism as the target of selection and the unit of evolution, the genetic information embodied by each of our microbiomes may itself be the target for and the product of the evolutionary process.
The conventional view, gleaned from studying macroscopic ecosystems, posits a sharp distinction between ecological and evolutionary timescales. When we speak of tropical rainforests or the Pacific Northwest intertidal zone, environmental changes occurring over the lifetime of an individual can elicit a physiological response; certain organisms can alter the expression of traits in response to sudden changes. In contrast, perturbations occurring over ecological time elicit responses in species composition and abundance; those stretching over larger timescales alter the genetic compositions of populations.
Ongoing climate change, for instance, highlights the difference between ecological and evolutionary timescales: Anthropogenically driven climate change is occurring too quickly to enable the majority of plant and animal species to respond through genotypic change. Instead, most species can only change their geographic distribution, a frequently foreclosed response to environmental disruption.
This distinction between ecological and evolutionary timescales appears fundamental, but may not apply when dealing with the microbiome. For many if not all members of the human microbial fauna, generation times are measured in hours or even minutes. These short generation times, coupled with the large population sizes of many bacteria, effectively elide the boundary between ecological and evolutionary time (this attribute also accounts for the fiendish ability of viruses to outrace both the immune system and efforts to combat viral infections).
Because one human lifetime may encompass a million bacterial generations, individual species and the microbiome itself can evolve within a single host. Bacteria can respond to changes in their environment with a seamless arsenal that ranges from transcriptional regulation to the rapid spread of an advantageous mutation. Bound as humans’ perceptions are by our own experience, we have trouble imagining the evolutionary changes unfolding within us. But for our bacterial partners, a human lifetime is deep time.
As scientists redefine the notion of biological identity, we are realizing that each person’s fate, from early in development to the grave, is inextricably linked to his or her microbiome. Evidence for this intertwining is now coming from many quarters. Diseases and syndromes that the medical profession never imagined would have a bacterial component, including psoriasis, asthma, colitis, obesity, and cardiovascular disease, have now been clearly associated with specific characteristics of the human microbiome. At the same time, biologists now realize that such attributes in some ways reflect the genome of the host: The gut microbiomes of identical twins differ by more than 50 percent of their component species, but are nonetheless more similar than those of fraternal twins. The extent to which each person’s microbiome is integral to survival, growth, and, aging is becoming increasingly clear.
The image of bacteria lying in wait to do people harm or, at best, hitching a free ride on our precious bodily fluids has been replaced by a far more interesting portrait of bacterial cells in constant and vital interaction with the eukaryotic cells in our bodies. These new, no-longer-invisible partners challenge human notions of identity and of the boundaries that define us.
A revolutionary notion of humans as superorganisms is now emerging. In this formulation, each person is, on average, an assemblage of 37 trillion eukaryotic cells combined with 300 trillion bacterial cells; the 20,000 protein-coding genes in the eukaryotic genome supplemented by 2 million bacterial genes.
Seen this way, much of modern medicine will need radical revision. Broad-spectrum antibiotics will have to give way to narrow-spectrum, targeted therapies. The root causes of many diseases will require a consideration of the state of the microbiome. Pathogenesis itself will need reframing as a consequence of disrupted microbial communities.
These complex and dynamic ecosystems, so inextricably linked to our lives, have forced microbiologists and physicians to revisit principles of community ecology established over the last century. Ecology has always combined fieldwork, experiment, and theory, and the microbiome is ripe for these approaches. The discovery of these complex and critical communities—literally living under our noses—provides ecology with new stomping grounds. As a result, scientific communities that rarely intersected—physicans and community ecologists, microbiologists and theoretical ecologists—are now poring over the same data.
These are early and exciting days, but one conclusion is clear: Mathematics and physics may thrive on rules that are independent of scale, but in biology the rules that govern the very small and short-lived may be different. We humans are among the largest and longest-lived of organisms, but we are not alone, nor, in the end, are we the measure of all things.
Editor's Note: This is a corrected version. For more information about the correction, please see: Royal Society Misquoted.
Click "American Scientist" to access home page
American Scientist Comments and Discussion
To discuss our articles or comment on them, please share them and tag American Scientist on social media platforms. Here are links to our profiles on Twitter, Facebook, and LinkedIn.
If we re-share your post, we will moderate comments/discussion following our comments policy.